In Focus
Mpox virus Clade Ib continues to be a closely monitored public health threat
Since November 2023, the Eastern parts of the Democratic Republic of the Congo (DRC) continue to see cases of the mpox virus Clade Ib that is also spreading through regional travel to neighboring countries in Central and Eastern Africa, incl. Burundi, Ethiopia, Kenya, Rwanda and Uganda. Outside of Africa, cases continue to be reported, including in Australia, Austria, Brazil, Canada, China, France, Germany, India, Israel, Italy, Japan, Netherlands, Oman, Pakistan, Portugal, Scotland, Slovakia, Spain, Sweden, Thailand, United Kingdom and the United States.
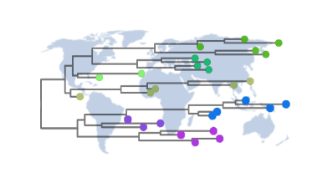
GISAID EpiPox provides access to the latest genome sequences and associated metadata of the mpox viruses, including Clade Ib and recombinant Ib+IIb. On 5 January 2026, Spain's Instituto de Salud Carlos III submitted a genome sequence of Clade Ib from a human case with no travel history. Previously, in the UK, a new recombinant (Ib+IIb) mpox was identified in a specimen sample collected from a traveller returning from Asia. The phylodynamics of Clade Ib can also be monitored on GISAID’s up-to-date phylogenetic trees.
An Equitable Pandemic Accord?
A perspective from a Congolese Researcher
Brazzaville, - Prof. Dr. Francine Ntoumi reflects on her surveillance work in Africa during COVID-19 and shares her views on global health equity, lessons learned, the impact of technology, and the importance of international collaboration and role of GISAID's Regional Hub in Central Africa.
Dr. Francine concludes her perspective with a cautionary reminder of what a global treaty will need to provide. See also: "Data imperialism in a pandemic." On 20. May 2025, the WHO Pandemic Accord was adopted by Member States at the Seventy-eighth World Health Assembly.
Upholding Trust: Inside GISAID’s Decision to Withdraw Data Feeds
During the COVID-19 response, some users with valid GISAID access credentials requested additional permissions, including data feeds to enable, e.g. dashboards to track variant prevalence or visualize the genomic evolution of SARS-CoV-2 across the globe. These requests were granted under supplementary agreements that required strict adherence to GISAID’s terms and assurances against misuse.
Subsequent audits, conducted with external experts and guided by GISAID’s Compliance Board, identified irregularities and breach of agreements which encroached upon contributors’ rights. In such cases, GISAID terminated the data feeds and revoked the supplementary permissions, as usage had diverged from the stated, approved purposes. Their regular access to GISAID, however, remained unaffected.
Despite this, a small but vocal group has circulated claims that access was cut “without justification,” mischaracterizing the circumstances. To uphold transparency and protect its community, GISAID is clarifying the record and sharing representative examples of cases where data feeds were withdrawn due to non-compliance. Read the full statement
H5N1 Bird Flu continues to circulate in the United States
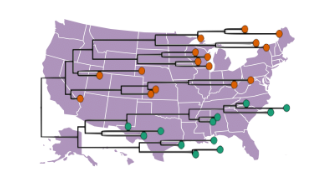
Clade 2.3.4.4b of the highly pathogenic avian influenza (HPAI) virus causing outbreaks in wild and domestic birds around the world continues to spread in dairy cows, poultry and other animals across the United States. Since April 2024, the U.S. CDC confirmed 70 human cases through genome sequence analysis. Data in some of these cases showed the amino acid substitution NA-S247N known to slightly reduce susceptibility to the neuraminidase inhibitor oseltamivir in laboratory tests. In one case, a different change in the polymerase acidic (PA) protein was detected.
On 19 March 2025, the CDC released specimen data from the first human case in Ohio. The specimen sequence belongs to genotype D1.3. Previously the CDC released data from the first human fatality involving a patient from Louisiana that had been exposed to non-commercial backyard poultry and wild birds.

CBD Study finds 'GISAID might provide useful lessons and insight to ABS discussions'
A peer-reviewed fact-finding and scoping study on digital sequence information on genetic resources in the context of the Convention on Biological Diversity and the Nagoya Protocol, highlights key advantages of GISAID’s sharing mechanism and a fair and equitable benefit-sharing resulting from access to data.
With the core principals of timely international sharing of health data for protecting populations against lethal infectious disease outbreaks and adherence to scientific etiquette of acknowledgement of the source of data has resulted in global trust and confidence in GISAID.
Phylodynamics
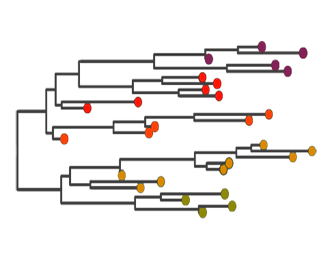
Phylodynamics
Tracking Variants
Frequency Dashboards
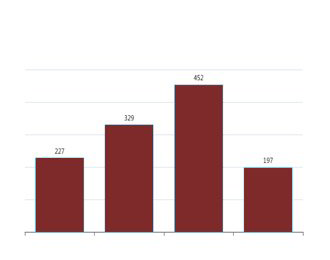
Variants and Frequencies
Public-Private Partnerships of the GISAID Initiative

The GISAID Initiative involves public-private partnerships, among them the partnership with the Federal Republic of Germany, and governmental public-health and academic institutions in Argentina, Brazil, China, Republic of the Congo, Ethiopia, Indonesia, Malaysia, Russia, Senegal, Singapore, South Africa, and the support by Friends of GISAID, a registered non-profit association and administrative arm of the Initiative.
Congratulations to GISAID for ten years of successful work on pandemic influenza preparedness. As one of the key players in ensuring effective data sharing GISAID has made a significant contribution to global health security
Prof. Jane Halton AO PSM
Chair, Coalition for Epidemic Preparedness Innovations CEPI
Commemorating the centenary of the 1918 pandemic, the most catastrophic event in the recorded history of influenza, it is reassuring to know that GISAID is ready and prepared when a similar event emerges and threatens global health. Congratulations on bringing together one of the most successful global collaborations ever achieved
Prof. Dr Rob Webster
St Jude Children’s Research
Hospital, Memphis, Tennessee
The unique contribution of the GISAID data sharing mechanism is the confidence it has engendered among scientific and political communities as it has added to their capabilities to collaborate more effectively to combat influenza viruses
Dr med David Nabarro
United Nations System Coordinat.
for Avian & Human Influenza (ret)
We do need substantially innovative mechanisms for microbe sharing, if mankind is to survive future pandemics. GISAID is an excellent example!!!
Dr Suwit Wibulpolprasert
Ministry of Public Health, Thailand
International Health Policy Program Foundation
IFPMA acknowledges GISAID’s important role in providing the platform for the open and timely sharing of influenza data and building greater trust among countries and stakeholders, a key element to influenza global pandemic preparedness
Thomas B. Cueni
Director General
International Federation of Pharma
Manufacturers & Associations
The tenth anniversary of GISAID represents a landmark in global solidarity. A pandemic strain of influenza is perhaps the world's greatest threat. Everything GISAID stands for: virus sharing, cutting-edge research, open access, and international cooperation to guarantee health security couldn't be more important
Prof. Lawrence O. Gostin
WHO Collaborating Center on
National and Global Health Law
Georgetown University
GISAID’s trustworthy data sharing principles forever transformed global collaboration in the fight against influenza, enabling unprecedented rapid response to outbreaks. In 2013, Nature called China’s sharing of H7N9 avian influenza data through GISAID ‘next to exemplary’
Prof. Dr George Fu Gao
Director General
Chinese Center for Disease Control and Prevention
The core principles enshrined in GISAID's sharing mechanism are a shining beacon of hope. A recognized leader in influenza pandemic preparedness, thanks to its brilliant performance in data sharing
Ambassador Makarim Wibisono
Republic of Indonesia (ret)
Professor, Universitas Airlangga
A key to protecting the world from future viral threats is having immediate and open access to critical viral data. GISAID has established a highly effective, trusted and time-tested model for influenza data sharing that could serve as an important model for other viral families
Dr Dennis Carroll
Global Health Security
US Agency for International Development (USAID)
GISAID successfully built upon the collaborative ethos of the 70-year old WHO Global Influenza Programme, to complement and extend the sharing of viruses, reagents & essential information
Dr med Wenqing Zhang
World Health Organization
Global Influenza Programme
The GISAID Initiative was established to champion (and enhance) rapid sequence data sharing for seasonal and pandemic influenza preparedness - a global public health imperative. GISAID’s success exceeded our expectations and provides an important model for rapid data sharing for other pathogens with pandemic potential
Dr Nancy J. Cox
WHO Collaborating Center for Surveillance, Epidemiology and Control of Influenza
Centers for Disease Control and Prevention (ret)
Not all big ideas become a reality and not all big ideas fill a global need. As a public-private partnership GISAID is a model for data sharing in the digital age. On its 10th anniversary we may look back at the initial inspiration and the headline of the supporting editorial in Nature that puts the point succinctly: Sharing saves lives
Dr med Bruce G. Gellin
Global Immunization, President
Sabin Vaccine Institute
ECDC congratulates GISAID for a successful 10 years of advocating for and implementing sharing of influenza sequence data. The initiative plays a key role in global and European pandemic preparedness and increases our understanding of the annual influenza seasons
Dr Mike Catchpole
European Centre for Disease
Prevention and Control (ECDC)
GISAID has advanced influenza virus data sharing to a new level, greatly contributing to the global effort to detect, respond, and mitigate seasonal and pandemic influenza
Prof. Dr med Peter Jay Hotez
Baylor College of Medicine, Dean
National School Tropical Medicine
I congratulate the GISAID Initiative on its ten-year anniversary as it continues its important work to promote the international sharing of influenza virus sequences and data
Ambassador John E. Lange
U.S. Department of State (ret)
United Nations Foundation
Over the past decade, GISAID has been an invaluable global partner in fostering open access to data related to influenza, a central issue related to influenza and all EIDs
Prof. Dr med Keiji Fukuda
The University of Hong Kong
School of Public Health
GISAID encourages increased collection and rapid dissemination of data that improves our understanding of the complex and dynamic epidemiology of influenza viruses. On behalf of OFFLU network, we offer our congratulations on the contribution GISAID has made to build international collaboration over the last 10 years
Dr Peter Daniels
Dr David Swayne
OFFLU OIE/FAO Network of
Expertise on Animal Influenza
The pioneering concept of transparent data sharing developed GISAID into the premier source of influenza virus sequence information and proven its worth in outbreak situations
Prof. Dr Thomas C. Mettenleiter
Friedrich-Loeffler-Institute
Federal Research Institute
for Animal Health, Germany
Ten years after GISAID first introduced its game-changing mechanism, breaking data sharing barriers, it continues to be a most trusted leader in pandemic preparedness & response
Prof. Dr Yuelong Shu
Sun Yat-sen University, Dean
School of Public Health, Shenzhen
GISAID has become the most complete public database for influenza virus sequence data in support of fundamental science and public and animal health applications
Prof. Dr Ron Fouchier
Erasmus MC Rotterdam
Viroscience & Nat'l Influenza Cntr
By sharing influenza virus sequences among scientists around the world, GISAID has had a tremendous impact on influenza virus research
Prof. Dr Yoshihiro Kawaoka
University of Wisconsin-Madison
University of Tokyo
From the latest seasonal to new zoonotic and animal influenza viruses, GISAID always has the most relevant strains and exciting new analysis tools
Dr Sebastian Maurer-Stroh
Agency for Science, Technology
and Research A*STAR Singapore
Data shared via EpiFlu, EpiCoV, EpiRSV, EpiPox, EpiArbo
22,323,801
genetic sequence submissions
GISAID Resources
![]()
Data Acknowledgement Locator